The CURVEFIT function uses a gradient-expansion algorithm to compute a non-linear least squares fit to a user-supplied function with an arbitrary number of parameters. The user-supplied function may be any non-linear function where the partial derivatives are known or can be approximated. Iterations are performed until the chi square changes by a specified amount, or until a maximum number of iterations have been performed.
This routine is written in the IDL language. Its source code can be found in the file curvefit.pro in the lib subdirectory of the IDL distribution.
Syntax
Result = CURVEFIT( X, Y, Weights, A [, Sigma] [, CHISQ=variable] [, /DOUBLE] [, FITA=vector] [, FUNCTION_NAME=string] [, ITER=variable] [, ITMAX=value] [, /NODERIVATIVE] [, STATUS={0 | 1 | 2}] [, TOL=value] [, YERROR=variable] )
Return Value
Returns a vector of values for the dependent variables, as fitted by the function fit. If A is double-precision or if the DOUBLE keyword is set, calculations are performed in double-precision arithmetic, otherwise they are performed in single-precision arithmetic.
Arguments
X
An n-element vector containing the independent variable values. X may be of type integer, floating point, or double-precision floating-point.
For multivariate models, X can also be an n x m or an m x n array, where n is the number of values and m is the number of independent variables. In this case your fit function needs to be written to correctly interpret the dimensions of X.
Y
A vector of dependent variables. Y must have the same number of elements as F returned by the user-defined function.
Weights
For instrumental (Gaussian) weighting, set Weightsi = 1.0/standard_deviation(Yi)2. For statistical (Poisson) weighting, Weightsi = 1.0/Yi. For no weighting, set Weightsi = 1.0. If Weights is set to an undefined variable then no weighting will be used. Set Weightsi to 0 to not include it in the fit.
A
A vector with as many elements as the number of terms in the user-supplied function, containing the initial estimate for each parameter. On return, the vector A contains the fitted model parameters.
Sigma
A named variable that will contain a vector of standard deviations for the elements of the output vector A.
Note: If Weights is omitted, then you are assuming that your supplied model is the correct model for your data, and therefore, no independent goodness-of-fit test is possible. In this case, the values returned for the Sigma argument are multiplied by SQRT(CHISQ/(N-M)), where N is the number of points in X, and M is the number of coefficients. See Section 15.2 of Numerical Recipes in C (Second Edition) for details.
Keywords
CHISQ
Set this keyword to a named variable that will contain the value of the chi-square goodness-of-fit statistic, weighted by the measurement error:
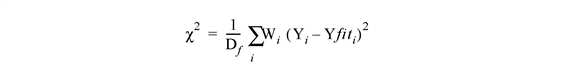
where Df = N - M is the degrees of freedom (N is the number of points in X, and M is the number of coefficients), W is the weighting value, and Yfit is the result.
DOUBLE
Set this keyword to force the computation to be done in double-precision arithmetic.
FITA
Set this keyword to a vector, with as many elements as A, which contains a zero for each fixed parameter, and a non-zero value for elements of A to fit. If not supplied, all parameters are taken to be non-fixed.
Note: If the NODERIVATIVE keyword is set then CURVEFIT will compute the partial derivatives by calling your function once for each parameter, using a value of that parameter that has been incremented by a small epsilon. This call will be made for each parameter including the fixed parameters. During the actual fitting procedure, IDL will ensure that the fixed parameters remain constant.
FUNCTION_NAME
Use this keyword to specify the name of the function to fit. If this keyword is omitted, CURVEFIT assumes that the IDL procedure FUNCT is to be used. If FUNCT is not already compiled, IDL compiles the function from the file funct.pro, located in the lib subdirectory of the IDL distribution. FUNCT evaluates the sum of a Gaussian and a second-order polynomial.
The function to be fit must be written as an IDL procedure and compiled prior to calling CURVEFIT. The procedure must accept values of X (the independent variable), and A (the fitted function’s initial parameter values). It must return values for F (the function’s value at X), and optionally PDER (a 2D array of partial derivatives).
The return value for F must have the same number of elements as Y. The return value for PDER (if supplied) must be a 2D array with dimensions
[N_ELEMENTS(Y), N_ELEMENTS(A)].
See the Example section below for an example function.
ITER
Set this keyword equal to a named variable that will contain the actual number of iterations performed.
ITMAX
Set this keyword to specify the maximum number of iterations. The default value is 20.
NODERIVATIVE
If this keyword is set, the routine specified by the FUNCTION_NAME keyword will not be requested to provide partial derivatives. The partial derivatives will be estimated by CURVEFIT using forward differences. If analytical derivatives are available they should always be used.
STATUS
Set this keyword to a named variable that will contain an integer indicating the status of the computation. Possible return values are:
|
0 |
The computation was successful.
|
|
1 |
The computation failed. Chi-square was increasing without bounds.
|
|
2 |
The computation failed to converge in ITMAX iterations.
|
TOL
Use this keyword to specify the desired convergence tolerance. The routine returns when the relative decrease in chi-squared is less than TOL in one iteration. The default value is 1.0 x 10-3.
YERROR
Set this keyword to a named variable that will contain the standard error between YFIT and Y.
Examples
Fit a function of the form F(x) = a * exp(b*x) + c to sample pairs contained in arrays X and Y. The partial derivatives are easily computed symbolically:
df/da = EXP(b*x)
df/db = a * x * EXP(b*x)
df/dc = 1.0
First, define a procedure to return F(x) and the partial derivatives, given X. Note that A is an array containing the values a, b, and c.
PRO gfunct, X, A, F, pder
bx = EXP(A[1] * X)
F = A[0] * bx + A[2]
IF N_PARAMS() GE 4 THEN $
pder = [[bx], [A[0] * X * bx], [replicate(1.0, N_ELEMENTS(X))]]
END
Compute the fit to the function we have just defined. First, define the independent and dependent variables:
X = FLOAT(INDGEN(10))
Y = [12.0, 11.0, 10.2, 9.4, 8.7, 8.1, 7.5, 6.9, 6.5, 6.1]
weights = 1.0/Y
A = [10.0,-0.1,2.0]
yfit = CURVEFIT(X, Y, weights, A, SIGMA, FUNCTION_NAME='gfunct')
PRINT, 'Function parameters: ', A
IDL prints:
Function parameters: 9.91120 -0.100883 2.07773
Thus, the function that best fits the data is:
f (x) = 9.91120(e-0.100883x) + 2.07773
Here is another example for CURVEFIT, which shows how to use multivariate input. The model function is:
Z = A X^B + C X Y + D Y^E
Click the following program:
PRO mymulticurvefit, XY, A, F, PDER
X = XY[*,0]
Y = XY[*,1]
F = A[0]*X^A[1] + A[2]*X*Y + A[3]*Y^A[4]
PDER = [[X^A[1]], [A[0]*X^A[1]*ALOG(X)], $
[X*Y], [Y^A[4]], [A[3]*Y^A[4]*ALOG(Y)]]
END
Then click to run the following commands on the command line to show the plot and overplot output:
seed = 1
n = 1000
X = 2*RANDOMU(seed, n)
Y = 2*RANDOMU(seed, n)
XY = [[X], [Y]]
A = [-3, 2, 5, 3, 3]
noise = RANDOMN(seed, n)
Z = A[0]*X^A[1] + A[2]*X*Y + A[3]*Y^A[4] + noise
A = [1d, 1, 1, 1, 1]
yfit = CURVEFIT(XY, Z, weights, A, $
FUNCTION_NAME='mymulticurvefit')
PRINT, 'Function parameters: ', A
iPlot, X, Y, Z, /SCATTER
X1 = 2*FINDGEN(101)/100
X = REBIN(X1, 101, 101)
Y = TRANSPOSE(X)
Z = A[0]*X^A[1] + A[2]*X*Y + A[3]*Y^A[4]
iSurface, Z, X1, X1, /OVERPLOT
Version History
|
Pre 4.0 |
Introduced |
|
5.6 |
Added YERROR keyword
|
|
6.0 |
Added FITA and STATUS keywords
|
See Also
COMFIT, GAUSS2DFIT, GAUSSFIT, LMFIT, POLY_FIT, REGRESS, SFIT, SVDFIT